1. How long will my antibody last if it is properly stored?
Our antibodies are guaranteed for 1 year after purchase. However, our internal studies show that the antibody will remain functional for a minimum of 5 years if properly stored.
2. How should I store my antibodies? Do I need to store it in aliquots?
Please refer to the "Storage Buffer" and "Storage Condition" sections on the product webpage or datasheet to obtain the storage buffer information and storage instructions for your specific product.
For antibodies that we store in a glycerol-based storage buffer, please store them in its original tube at -20°C. These products are stored in 50% glycerol, which prevents them from freezing. Aliquoting is unnecessary for -20°C storage and is not recommended.
Our PBS Only antibodies are stored at -80°C. Aliquoting is recommended for these products to avoid multiple freeze-thaw cycles.
Generally, our conjugated and unconjugated primary antibodies designed and validated for use in flow cytometry are stored at 4°C.
3. Where can I find detailed information about an antibody?
On our antibody product pages we provide information on the antibody isotype, tested applications and species, recommended antibody dilutions, protocols and publications which the antibody is cited in. Validation data from our in-house tests are displayed on the product page, with corresponding protocol details found in our product-specific protocols. Datasheets are available to download in PDF format on each product page. If you require any further information, please contact us and we'd be happy to help.
4. Where can I find the concentration of my antibody?
The concentration of a given antibody is found on the product datasheet and on the product vial tube itself. In the event that the concentration displayed on the vial tube is different to that shown on the product datasheet, please refer to the vial tube concentration. We previously measured our antibody concentration by Bradford assay, and have now switched to Nanodrop measurement. You can find out more about the difference in this blog.
5. Where can I find the lot number of my antibody?
The lot number of your antibody can be found under the barcode on the vial. If you have any issue locating the lot number, please contact us and we can assist you with this.
6. How can I find the immunogen sequence of an antibody?
Under the product information section of the product page, there is a section which lists the immunogen (Figure 1). Clicking this link will take you to the product page for the fusion protein immunogen, which lists the amino acid sequence (Figure 2). Should you require more detailed information, please contact us.
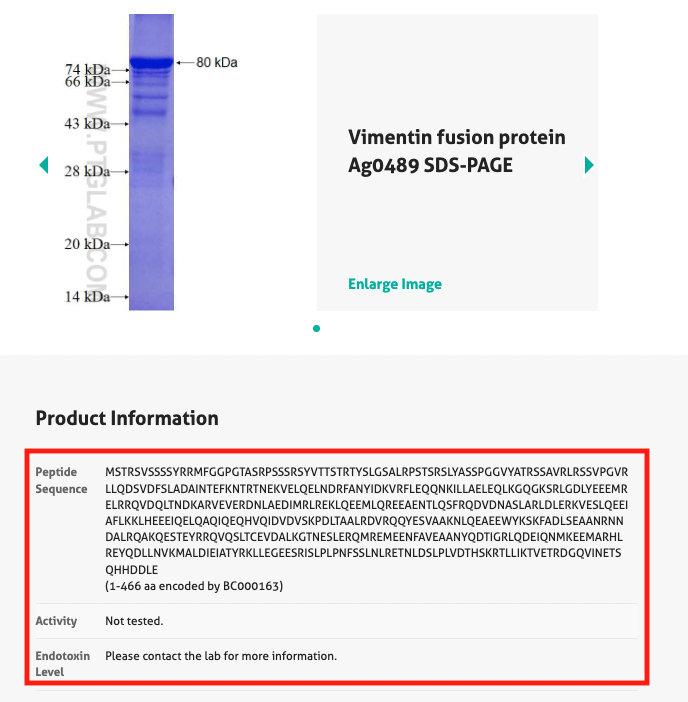
7. Is there a blocking peptide available for my antibody?
The fusion proteins used as the immunogen in the production of an antibody can be used as a blocking peptide. You can find the instructions on where to find this in Section 5 above (Figure 1).
8. Has my antibody been tested on any species other than human?
We list the species that we have validated the antibody in on our product pages under 'Tested Reactivity'. In addition, we also list the species in which the antibody has been used in publications which cite it and list this under 'Cited reactivity'. Under our Proteintech guarantee, we cover you when using our antibodies in any species.
9. Which applications can my antibody be used in?
All currently tested or cited applications are listed on each product page. Under our Proteintech guarantee, we cover you when using our antibodies in any application.
10. Does my antibody cross-react with other species?
Cross-reactivity depends on the extent of protein sequence similarity between the immunogen and the potential cross-reactive protein sequence. A pair-wise sequence alignment can be performed online through the NCBI-BLAST website. If you require any further information about potential cross-reactivity, please contact us.
11. Has the antibody been cited in any publications?
Each product page shows the number of publications and has links to the publication if the antibody has already been cited. The publication list can be found at the bottom of the product page and allows you to filter the publications by application, date and journal impact factor (Figure 3).
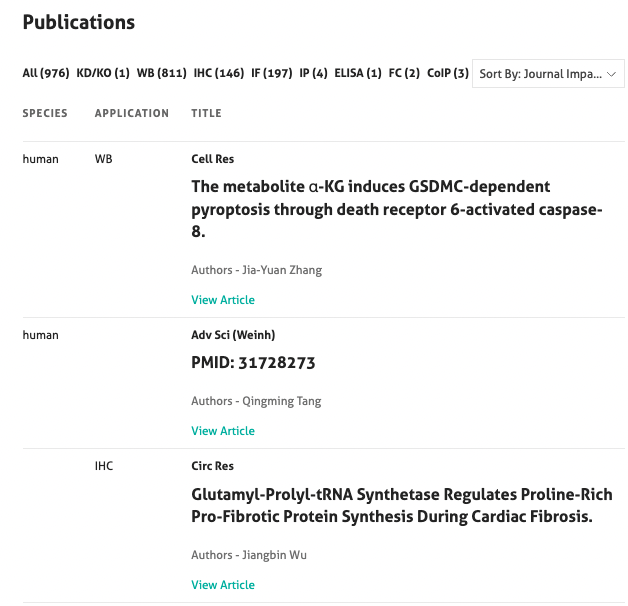
12. What is the clone number?
Each monoclonal antibody has a clone number that is specific for its single clone of hybridoma cells. The clone number can be found on the product page next to the catalog number (Figure 5).

13. How should I choose a suitable secondary antibody?
The choice of appropriate secondary antibody is determined by the specifications of the primary antibody and the application (i.e., species, subclass, and fragment type).
Guiding principles for choosing a suitable secondary antibody:
- The secondary antibody must be raised against the host species of the primary antibody (e.g., anti-mouse or rabbit)
- The secondary antibody must be specific to the isotype class of the primary antibody (e.g., IgG or IgM)
- If the primary antibody is a fragment (e.g., F(ab’)2), the secondary antibody should target that specific fragment to reduce background noise
Proteintech offers a variety of secondary antibodies suitable for Western Blotting, ELISA, cellular imaging, and flow cytometry. View our secondary antibodies blog for more detailed information.
14. How should I choose an isotype control?
Isotype controls are used to determine non-specific interactions in a given sample such as Fc receptor binding. Most isotype controls are monoclonal and most are not suitable for use with polyclonal antibodies as they contain more than one IgG class. The isotype control antibody should match the primary antibody regarding host species, isotype, and conjugation. Lastly, the sample incubated with the isotype control antibody and the sample incubated with the primary antibody should always be run in parallel.
15. How can I choose a positive and a negative control?
A sample without a primary antibody can be used as a negative control to distinguish specific and non-specific signal from the secondary antibody. Any products/stainings obtained with these control conditions can be attributed to non-specific (off-target) interactions. Furthermore, samples that have been genetically manipulated to alter the target’s levels (e.g., CRISPR) can be used as a negative control.
For a positive control, any tissues, cells, or lysates that have been used successfully in our validation data set or by our customers are a suitable control. Each antibody has its own PubMed publication record on our website. Additionally, the Uniprot, Omnigene, and GeneCards databases are great resources for finding tissues/cell lines with high expression of the target. The latest publications record in PubMed can also be a useful tool for detailed and the most up-to-date information regarding target research.
16. Why is the predicted WB band size different from the actual one?
There may be several reasons for a difference in band size, such as multimers, gel migration, relative charges, splice variants, post-translational modification, or cleavage. For more information, please contact our technical support.
17. Should I perform an antigen retrieval step for my IHC experiment?
Yes, it is recommended to perform an antigen retrieval step since the fixation process during paraffinization cross-links proteins and may mask the epitopes, resulting in weak or false negative staining. However, this situation can be rectified with a heat-induced epitope retrieval (HIER) or proteolytic-induced epitope retrieval (PIER) step. The most suitable choice depends on the tissue type and primary antibody.
For more detailed information regarding antigen retrieval steps, please check our IHC guide and protocols page.
1. What method does Proteintech use for recombinant protein purification?
We usually add 6x His tags or GST tags to the N-terminus of proteins; these proteins are then purified by Ni2+ and glutathione affinity chromatography, respectively.
2. What storage buffers are used for your recombinant protein?
The His-tagged recombinant protein is usually stored in 1x PBS buffer (58mM Na2HPO4, 17mM NaH2PO4, 68mM NaCl, pH 7.4) containing 300mM imidazole.
The GST-tagged recombinant protein is usually stored in 1x PBS buffer (58mM Na2HPO4, 17mM NaH2PO4, 68mM NaCl, pH 7.4) containing 100mM glutathione (GSH).
1. What does the ELISA kit contain?
-
Antibody-coated 96-well microplate
-
Assay standard
-
Detection antibody
-
HRP-conjugated antibody
-
Sample diluent
-
Detection diluent
-
Wash buffer
-
Tetramethylbenzidine (TMB) substrate
-
Stop solution
-
Plate cover seals
2. Which ELISA type do you offer?
All Proteintech ELISA kits are two-site sandwich ELISAs.
3. Are the plates pre-coated?
Yes, all plates are pre-coated with an antibody that is specific for the target.
4. Do I have to run the ELISA standards and samples in duplicate/triplicate?
It is recommended to run the experiment in duplicate at a minimum. Triplicate would be ideal.
5. Is it possible to use the reagents of two different ELISA kits/lots?
No, different lot numbers cannot be mixed.
6. Are shorter or longer incubation times possible?
Incubation times should be followed exactly as stated in the manual.
7. How should I store the reagent?
Proteintech ELISA kits can be stored at 4C for 6 months after receipt.
8. Can I use a non-validated sample type with this ELISA kit?
Most of our ELISA kits are validated for cell culture supernatant, plasma, or serum.
9. How sensitive are the ELISA kits?
The ELISA sensitivity is highlighted in the data sheet for each kit.
10. Why does my standard shows very low or no color development?
There are multiple explanations: incorrect storage, incorrect incubation time, improper antibody, insufficient washing, etc. For more information, please contact technical support.
11. What are the reasons for high background?
Improper washing, contaminated reagents, samples, or wells. For more information, please contact technical support.
12. Is it possible to store samples after the collection?
Fresh samples are ideally recommended. If you need to keep the samples, you can store at 4 °C for short-term storage(<3 days) or at -20 °C/-80°C for long-term storage(up to 6 months). Please avoid freeze/thaw cycles.
1. How long will my custom production project take?
Custom production projects can typically take up to 6 months to complete, but due to the complexity of the process, we cannot always give a guarantee of an exact schedule. Please see below for the schedule of our 102-day immunization protocol, which demonstrates our commitment to making sure that you receive an antibody that works.
102-Day Immunization Protocol
| Day 0 Pre-Immune Bleed | Day 56 Test Bleed & ELISA | Day 88 Production Bleed |
| Day 1 Primary Injection | Day 60 Boost 3 | Day 102 Final Bleed |
| Day 28 Boost 1 | Day 74 Production Bleed | |
| Day 42 Boost 2 | Day 78 Boost 4 |
2. Can I express the fusion protein in my lab and send you that?
No, unfortunately we do not currently accept fusion proteins expressed by our customers.
3. What carrier proteins do you use?
KLH is the default carrier protein. This is linked to the peptide via a terminal cysteine (added by Proteintech®). If you have cysteine residue in the middle of your peptide then we can use MAP beads instead.
4. How many rabbits are immunized for each peptide service?
Each antigen will be injected into two rabbits.
5. Should I choose a full-length fusion protein or peptide antigen?
In our experience, full-length fusion protein antigens usually generate antibodies with higher titers and higher sensitivities. We recommend the full-length fusion protein procedure if your protein of interest has low sequence similarity with other proteins. However, if you need an antibody that is specific to a certain protein, the sequence of which is highly similar to other proteins, we strongly recommend using a peptide antigen. Other reasons you might choose a peptide antigen include:
1. Not having the cDNA for the protein of interest;
2. Expression of the full-length fusion protein has not been successful for some reason.
6. Is your custom antibody production service limited to human genes?
No, we carry out custom production of antibodies for proteins from any species, depending on antigenicity.
7. How much antiserum will I receive?
Each rabbit will yield approximately 12 ml of serum in each production bleed and 45–50 ml of serum from 90–100 ml of blood in the final bleed, totalling about 150 ml of antiserum from two rabbits.
8. How does Proteintech determine which rabbit to purify serum from?
We decide that based on the ELISA results with the test bleeds. If both rabbits give good results, we will purify 15 ml of serum from each rabbit. If one rabbit gives better results than the other, we will purify 30 ml of serum from this rabbit.
9. How much purified antibody can I get from the antiserum?
It depends on the concentration of the specific antibody in the serum provided. Usually we can get 2–10 ml of purified antibody from 15 ml of antiserum.
1. What is the range for EC50?
The EC50 is defined as the concentration at which the bioassay activity is 50% of the maximum response.
2. What are the advantages of Humankine products over E. coli, CHO, and insect cell expressed proteins?
Recombinant Humankine cytokines and growth factors are expressed in an optimized, proprietary human cell line of HEK293 origin. Proteins produced using our technology have native glycosylation, folding, dimerization, and post-translational processing, so their biological activity and stability are superior to proteins expressed in non-human systems.
3. How are Humankine products shipped?
Humankine products are shipped at room temperature (RT) as lyophilized and carrier-free proteins. Before opening, we recommend briefly centrifuging the vial and then proceeding to follow the product-specific instructions as described either on the datasheet and/or certificate of analysis to reconstitute.
4. How should I store Humankine products?
Proteins are stable for one year from the date of receipt if stored between (-20°C) and (-80°C). All products are lyophilized, except for Thrombin (HZ-3010).
5. In what applications can Humankine products be used?
Humankine products are broadly used for stem cell research and cell therapy. At the time of writing, Humankine products have been cited in more than 250 PubMed publications. See the specific product pages for more details.
6. What type of media does Proteintech use for Humankine production with our expression system? Does Proteintech use animal components such as FBS?
We use commercially available, chemically defined serum and xeno-free media. It is completely free of animal components.
7. What is the recommended concentration for the reconstitution step?
We recommend reconstituting our proteins at less than or equal to 1mg/mL (except Wnt3a).
8. Is your cell line available from ATCC?
Should you require more details or information about our cell line, please contact our team.
9. Does Proteintech manufacture a GMP-grade recombinant protein?
Please contact us for more information on GMP-grade proteins. GMP proteins can be produced on custom request.
View FAQs for FlexAble Antibody Labeling Kits here.
Nanobodies/VHH
1. Are ChromoTek Nanobodies monoclonal or polyclonal?
ChromoTek Nanobodies are monoclonal and are recombinantly produced.
2. Do the Nanobodies have any tags?
Yes, most of the ChromoTek Nanobodies have a C-terminal 6x His-tag.
3. Do ChromoTek Nanobodies/VHHs bind to protein A or protein G?
Some Nanobodies bind to protein A, but not to protein G.
Nano-Traps
1. How much cell extract/total protein amount should I use for an immunoprecipitation reaction with Nano-Traps?
We recommend using 0.5 - 1.0 mg of total protein for one immunoprecipitation reaction. The amount of protein is also dependent on the expression level of your protein of interest and its interaction partners.
2. How many mammalian cells are required for an immunoprecipitation reaction with Nano-Traps?
For one immunoprecipitation reaction, we recommend using ~10^6 - 10^7 mammalian cells. The amount of cells is also dependent on the expression level of your protein of interest and its interaction partners.
3. What is the binding capacity of a Nano-Trap?
The binding capacity of each ChromoTek Nano-Trap and individual matrix version differs due to the conjugation efficiency of the Nanobody to the beads. However, regardless of the Nano-Trap specificity and matrix, 25µL of bead slurry is sufficient for an immunoprecipitation reaction. Refer to the individual product manual or webpage for the specific binding capacity.
4. Should I use an N-terminal or C-terminal tagged protein of interest for immunoprecipitation with Nano-Traps? Can I also tag my protein internally?
Both N-terminal or C-terminal fusion tags work well with Nano-Traps. Often C-terminal fusion tags are preferred, since only fully translated proteins will be captured or detected when using a Nano-Trap. The compatibility of an internal tag depends on the fusion protein and must be tested case by case.
5. Are Nano-Traps compatible with detergents?
Yes, ChromoTek Nano-Traps are highly compatible with high concentrations of common detergents such as NP-40, Triton X-100 or SDS. Refer to the individual product manual for detailed compatibilities.
6. Are Nano-Traps stable in the presence of reducing agents?
Yes, ChromoTek Nano-Traps are highly stable at high concentrations of common reducing agents such as DTT, TCEP or beta-mercaptoethanol. Since the Nanobodies of the Nano-Traps don’t contain surface-exposed disulfide bridges, reducing agents do not impair the Nanobody’s structure or function. Refer to the individual product manual for detailed compatibilities.
7. Should I pre-clear my sample when using a Nano-Trap?
Pre-clearing is often not necessary since ChromoTek Nano-Traps have a very low background and very low unspecific binding to host cell proteins. However, if pre-clearing is needed, we recommend using Binding Control Agarose Beads (bab-20) or Binding Control Magnetic Agarose Beads (bmab-20). Since the Magnetic Particles M-270 matrix is inert, pre-clearing is not needed when using Nano-Traps Magnetic Particles M-270.
8. Is the Nanobody released from the beads during elution after immunoprecipitation with a Nano-Trap?
No, the Nanobody of ChromoTek Nano-Traps is covalently conjugated to the bead matrices. Therefore, the Nanobody is not released from the beads in the elution step after an IP experiment.
9. How can I elute bound proteins from Nano-Traps under native conditions?
Most of the ChromoTek Nano-Traps can be eluted with 0.2 M glycine pH 2.5 at room temperature. Ensure to neutralize the eluate immediately after elution by adding 1 M Tris base pH 10.4. In addition, Nano-Traps binding to small epitope tags like V5-tag or Myc-tag can be eluted with the corresponding peptide.
10. Do I need to elute bound proteins from Nano-Traps for mass spectrometry analysis?
No, you can conduct an on-bead digestion directly after immunoprecipitation. This procedure allows faster and more efficient sample preparation and potentially higher yield.
11. Do I need to elute my protein of interest from Nano-Traps for enzymatic assays?
No, you can perform your enzymatic activity assay directly on the beads if the active center is not blocked.
12. Can I elute my protein of interest from Nano-Traps with chaotropic agents such as 8M Urea?
No, elution from ChromoTek Nano-Traps is not possible with chaotropic agents such as 8M Urea. The binding of the tagged protein to the Nano-Trap is highly stable preventing the quantitative release of the protein of interest in the presence of chaotropic agents.
GFP-Trap®
1. Is it possible to elute bound proteins from the GFP-Trap® with free GFP-Protein?
Elution with free GFP is not recommended. Due to the very high affinity of the GFP-Trap®, elution requires at least 400 µM of free GFP and isn't quantitative. The eluate then contains a very high amount of free GFP and a small amount of GFP-tagged fusion protein.
2. Does the GFP-Trap® bind TurboGFP?
No, the GFP-Trap® doesn't bind TurboGFP. TurboGFP is a green fluorescent protein derived from CopGFP of the copepod Pontellina plumata, which shares only ~20 % sequence identity with the commonly used GFP variants. If you want to pull down TurboGFP, the Chromotek TurboGFP-Trap should be used.
3. Why do I sometimes not see fluorescence after acid elution of my GFP-tagged protein from the GFP-Trap®?
Fluorescence is lost in some cases when the GFP-tagged protein is eluted with glycine buffer because the strong acidic pH 2.5 of glycine quenches the fluorescent signal of GFP. GFP fluorescence can be restored by neutralizing the eluate with 1 M Tris base pH 10.4 buffer immediately after elution.
DYKDDDDK Fab-Trap™
1. What is the amino acid sequence of the 1x and 3xFlag®-tag?
The amino acid sequence of 1xFlag®-tag is DYKDDDDK. The amino acid sequence of the 3x-Flag®-tag is DYKDHD-G-DYKDHD-I-DYKDDDDK. Note that the sequence of the 3xFlag®-tag is an optimized peptide sequence.
2. Does the DYKDDDDK Fab-Trap™ bind to 1x and 3xFlag®-tag?
Yes, DYKDDDDK Fab-Trap™ binds efficiently to both the 1x and 3xFlag®-tag. Elution of 1x and 3xFlag®-tagged proteins should be done with the 3xDYKDDDDK-peptide.
3. Should I use a N-terminal or C-terminal Flag®-tag for the DYKDDDDK Fab-Trap™? Can I also tag my protein internally?
Both N-terminal or C-terminal fusion tags work well with the DYKDDDDK Fab-Trap™. Often C-terminal fusions are preferred as only complete translation products are captured or detected when using an anti-Flag®-tag reagent. The compatibility of an internal DYKDDDDK-tag depends on the fusion protein and must be tested case by case.
4. What is the Fab-fragment of the DYKDDDDK Fab-Trap™ based on?
The DYKDDDDK Fab-Trap™ consists of a Fab-fragment derived from the mouse IgG1 clone M2, which is covalently bound to Agarose beads. It recognizes the same epitope as the M2 antibody. Due to its small size, the Fab-fragment has several advantages for IP, such as a lower background and broader wash buffer compatibility.
5. How can I avoid releasing the Fab-fragment from the DYKDDDDK Fab-Trap™ beads during elution such that it doesn't become visible on SDS-PAGE or Western Blot?
The Fab-fragment is released during elution with SDS-sample buffer and is visible on SDS-PAGE or WB at 25 kDa. To avoid the release of the Fab-fragment, you may use milder elution options such as elution with low pH or peptide elution with the 3xDYKDDDDK-peptide.
6. How can I detect the 1x or 3xFlag®-tagged protein by Western Blot?
To detect your Flag®-tagged protein by Western blot, we recommend using an anti-Flag®-tag antibody that is not from the host species mouse. The Fab-fragment of the DYKDDDDK Fab-Trap is from the host species mouse, and is released from the beads during elution with SDS-sample buffer. Using a mouse anti-Flag®-tag antibody (e.g. the M2 clone) will result in the detection of the 25 kDa Fab-fragment in your Western blot.
We recommend using a rabbit anti-Flag®-tag (e.g. Proteintech's DYKDDDDK tag polyclonal antibody, 20543-1-AP) in combination with an anti-rabbit secondary antibody. Alternatively, an HRP-conjugated anti-Flag® primary antibody can be used.
7. How can I perform pre-clearing of my sample before using the DYKDDDDK Fab-Trap™?
For pre-clearing of your sample, we recommend using our Binding Control Agarose Beads (bab-20). Pre-clearing with an isotype control antibody is not necessary since the DYKDDDDK Fab-Trap™ is only 50 kDa in size and lacks the Fc domain, which minimizes unwanted interactions with the host cell proteome.
8. What is the affinity (Kd) of the DYKDDDDK Fab-Trap™?
The dissociation constant Kd of the Fab-fragment of the DYKDDDDK Fab-Trap™ to a Flag®-tagged fusion protein is 36 nM.
9. Do I need to conjugate the DYKDDDDK Fab-Trap™ to protein A or G beads?
No, the DYKDDDDK Fab-Trap™ is a ready-to-use reagent. It consists of a Fab-fragment covalently conjugated to Agarose beads for convenient and time-efficient immunoprecipitation. Therefore, you do not need to conjugate it with protein A or G beads before performing an IP experiment.
Myc-Trap®
1. Does the Myc-Trap® bind endogenous c-Myc protein?
No, the Myc-Trap® does not bind to endogenous c-Myc protein under native conditions. Some epitope residues that have shown to be crucial for binding to the Myc-Trap® are buried in the three-dimensional structure of the c-Myc protein.
Nano-Booster/Nano-Label
1. How many dye molecules are coupled to Nano-Boosters and Nano-Labels?
Each Nano-Booster and Nano-Label molecule carries 1-2 fluorophores. Depending on the conjugated fluorophore, some Nano-Boosters are labeled in a site-directed way and carry either 1 or 2 dyes. Other Nano-Boosters and Nano-Labels are randomly labeled and carry on average 1-2 dye molecules.
2. Are Nano-Labels and Nano-Boosters applicable for live-cell imaging?
Nano-Labels and Nano-Boosters are small proteins (~15 kDa) and therefore are not able to penetrate through non-permeabilized cell membranes. With different protein transduction methods and reagents (e.g., electroporation), it is possible to introduce small amounts of Nano-Boosters and Nano-Labels into live cells. However, from our experience, the most efficient way is to microinject the Nano-Labels and Nano-Boosters.
3. Are Nano-Booster and Nano-Label suitable for Super-Resolution Microscopy?
Yes, Nano-Boosters and Nano-Labels are highly suitable for Super-Resolution Microscopy. Due to their small size of 2-3 nm, they minimize the linkage error and provide a more precise and dense staining than conventional antibodies (15 nm linear dimension). The selection of a Nano-Booster and Nano-Label conjugate depends on your microscope setup and lasers.
4. Can I do a simultaneous co-staining with two or more Nano-Boosters and Nano-Labels?
Yes, you can combine the Nano-Boosters and Nano-Labels. Therefore, you can mix Nano-Boosters and Nano-Labels in blocking solution and co-stain the sample simultaneously.
Spot Capture & Detection System
1. Should I use Spot-Trap® or Spot-Cap® for immunoprecipitation?
The Spot-Trap® is ideal for immunoprecipitation experiments because it has a very high binding affinity, is stable under harsh buffer conditions, and has minimal background signal. The Spot-tagged POI can be eluted directly using SDS sample buffer prior to detection via Western blotting.
2. Should I use Spot-Cap® or Spot-Trap® for protein purification?
The Spot-Cap® is well suited for protein purification because it has a high binding capacity, and the protein can be eluted under gentle conditions like 100 µM Spot-peptide or 100 mM glycine at pH 2.0 and +4 degrees. The gentle elution methods help to keep the native folding of your protein of interest which is beneficial for several downstream applications. Additionally, the Spot-Cap® is optimized for low contamination of host cell proteins.
3. Should I use an N-terminal or C-terminal Spot-fusion?
Both, N- or C-terminal fusions work well. Often C-terminal fusions are preferred as only complete translation products are captured or detected when using an anti-Spot-Tag® reagent.
4. Can I also insert the Spot-Tag® in the middle of my protein?
The use of the Spot-Tag® for internal protein tagging must be considered on a case-by-case basis. The Spot-Tag® should be present in a linear form and be accessible without steric hindrance from other parts of the protein of interest. Therefore, it is recommended to insert the Spot-Tag® into a sufficiently large and unstructured loop, an inherently unstructured domain, or a long domain linker. In addition, it is also important to ensure that the folding and structure of the protein of interest is not impaired by Spot-Tag® insertion.
5. Should I use a single Spot-Tag® or multiple Spot-Tag® repeats fused to my protein of interest?
Since the affinity between Spot-Tag® and the Spot Nanobody is very high, a single Spot-Tag® is sufficient for efficient detection in most cases. However, multiple Spot-Tag® repeats can increase the binding efficiency due to an avidity effect when immunoprecipitating with Spot-Trap®. When using multiple Spot-Tag® repeats, it is recommended to insert a longer linker sequence between the two tag sequences to allow multiple Spot Nanobodies to bind next to each other.
6. How can I detect my Spot-fusion in Western Blot application?
You can use ChromoTek's Spot-Tag® Antibody [28A5] to detect your Spot-tagged fusion protein. Alternatively, you can use ChromoTek's Spot-Label® for fluorescent WB detection.
Chromobodies®
1. When should I image my cells after transfection with the Chromobody® plasmid?
The Chromobody signal is maintained up to 3 days in the cell. However, this also depends strongly on the cell type. We generally recommend imaging the cells 16-24 hours after transfection.
2. Are Chromobodies® constitutively expressed?
Yes, Chromobody expression is regulated by immediate early promotor CMV. Therefore, Chromobodies are constitutively expressed, and their fluorescent signal can be observed within the first day after transfection.
3. Can the Chromobodies® diffuse through the cell membrane into the growth medium?
No, Chromobodies are small proteins expressed in the cytosol. They are not secreted into the medium and remain in the cell as long as the cell maintains its plasma membrane integrity.
4. Are Chromobodies® fluorogenic or do they only emit fluorescence when bound to a target?
Chromobodies are chimeric proteins consisting of a VHH fused to a fluorescent protein. They emit their fluorescence regardless of whether they are bound to a target or not.
5. Do Chromobodies only work in living cells?
Yes, the Chromobody is only expressed in living cells. Cells should be transfected with the Chromobody plasmid at least one day before imaging to observe the Chromobody location signal. However, cells can be fixed prior to imaging.
6. Can I propagate the Chromobody® plasmid in bacteria?
Yes, the Chromobody plasmids can be propagated in E.coli by standard techniques.
