Tested Applications
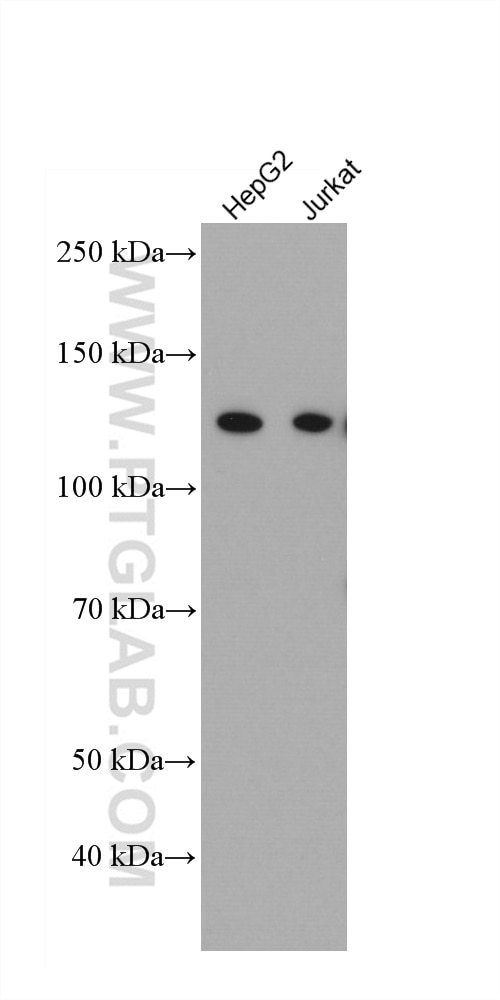
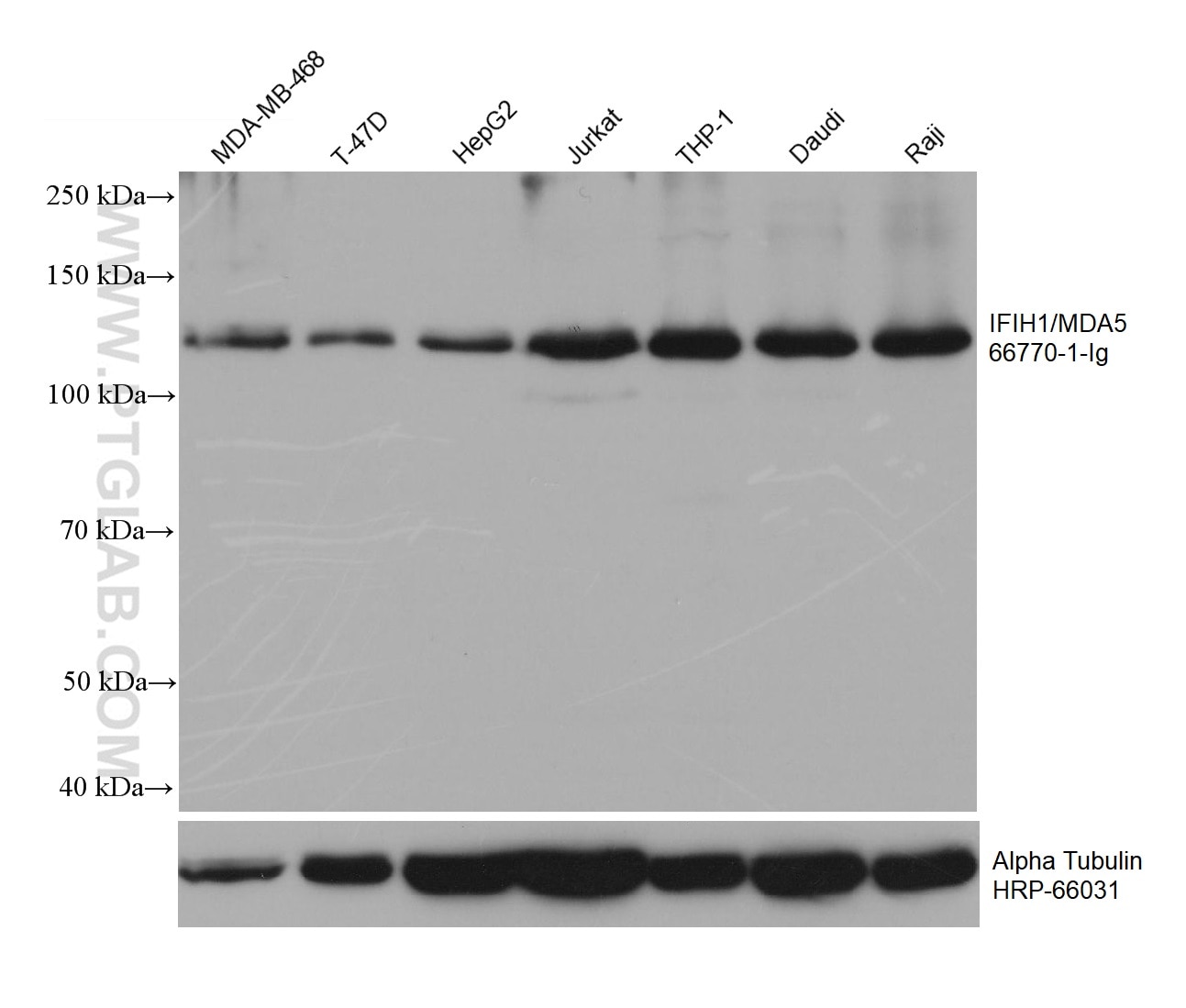
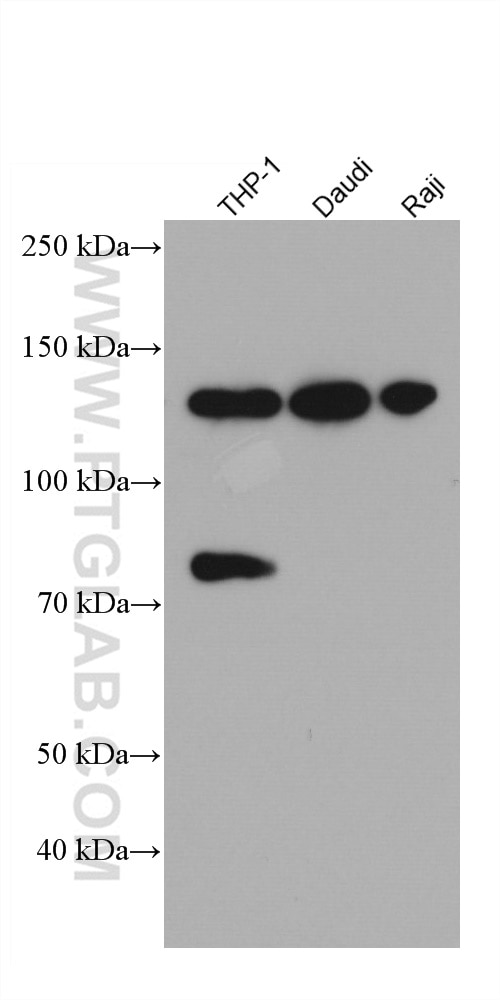
| Positive WB detected in | MDA-MB-468 cells, HepG2 cells, THP-1 cells, Jurkat cells, Daudi cells, Raji cells, T-47D cells |
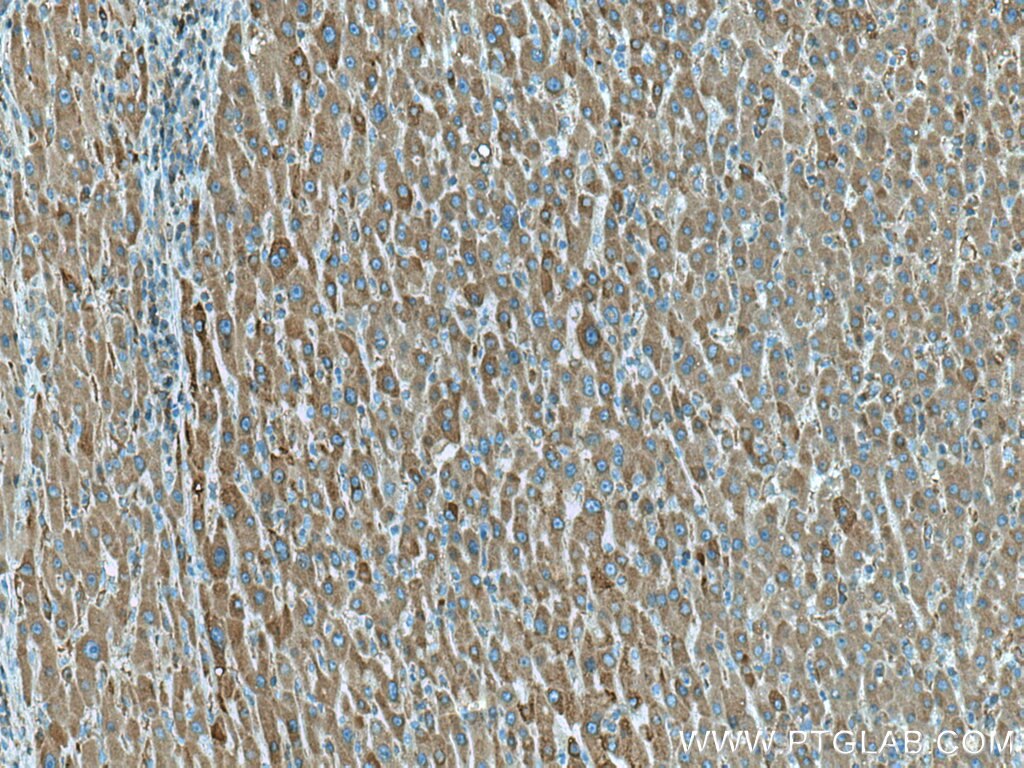
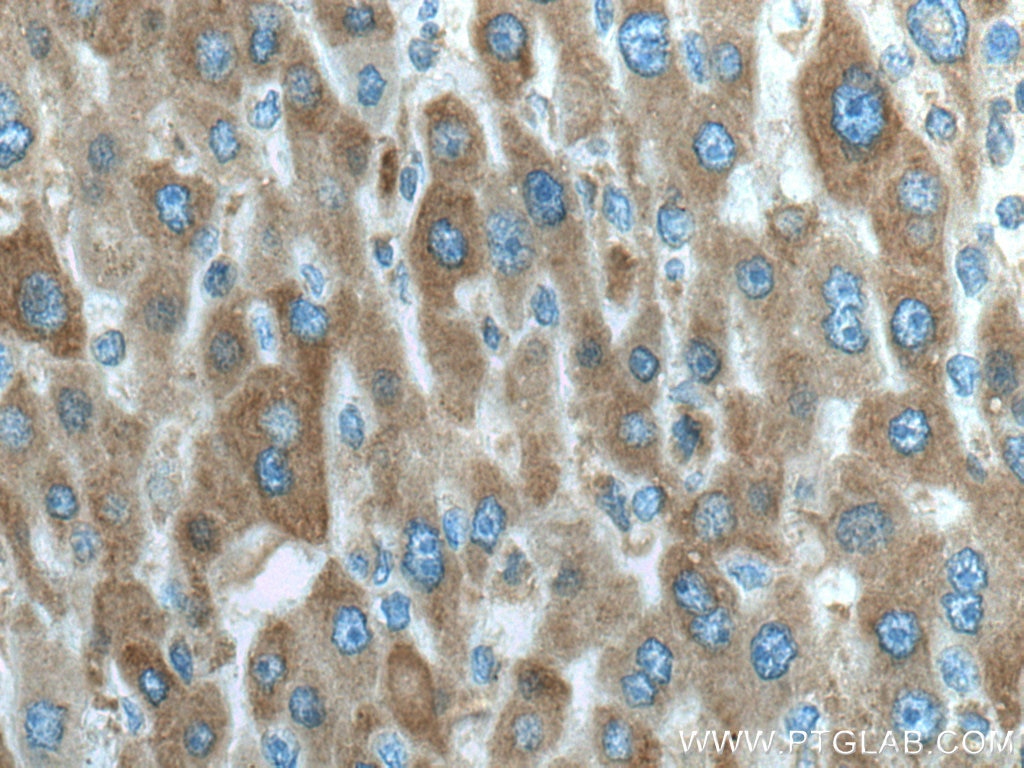
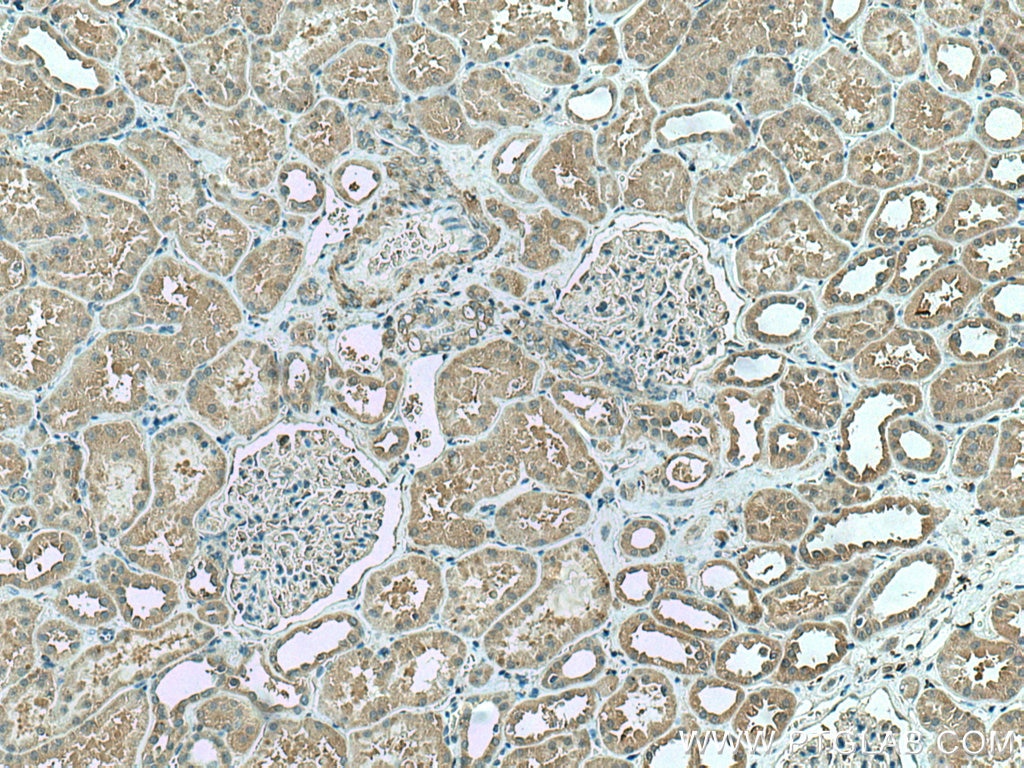
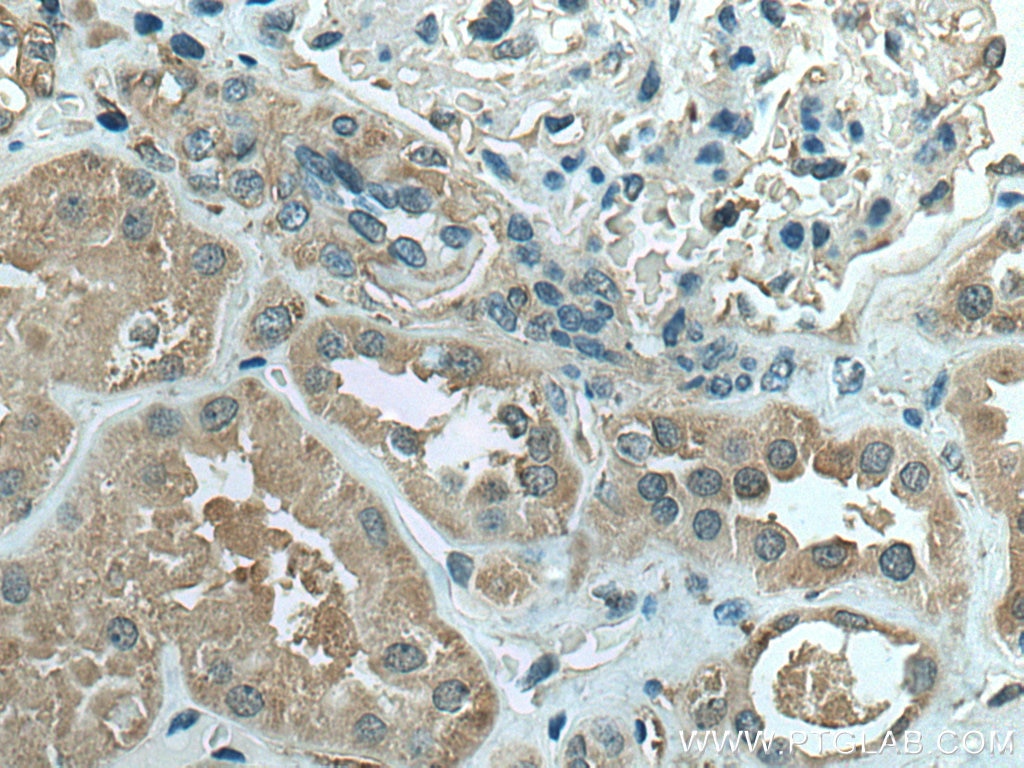
| Positive IHC detected in | human liver cancer tissue, human kidney tissue Note: suggested antigen retrieval with TE buffer pH 9.0; (*) Alternatively, antigen retrieval may be performed with citrate buffer pH 6.0 |
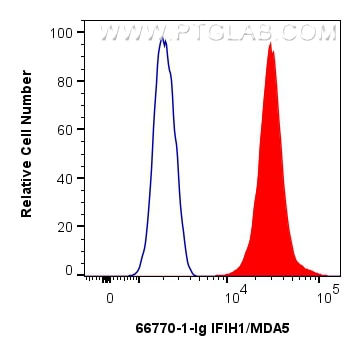
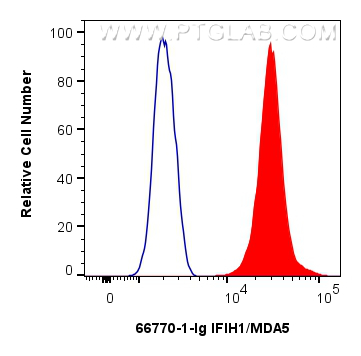
| Positive FC (Intra) detected in | Jurkat cells |
Recommended dilution
| Application | Dilution |
|---|---|
| Western Blot (WB) | WB : 1:5000-1:50000 |
| Immunohistochemistry (IHC) | IHC : 1:250-1:1000 |
| Flow Cytometry (FC) (INTRA) | FC (INTRA) : 0.25 ug per 10^6 cells in a 100 µl suspension |
| It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
Published Applications
| KD/KO | See 1 publications below |
| WB | See 8 publications below |
| IHC | See 1 publications below |
| IF | See 1 publications below |
Product Information
66770-1-Ig targets IFIH1/MDA5 in WB, IHC, IF, FC (Intra), ELISA applications and shows reactivity with human samples.
| Tested Reactivity | human |
| Cited Reactivity | human, mouse, pig, bovine |
| Host / Isotype | Mouse / IgG1 |
| Class | Monoclonal |
| Type | Antibody |
| Immunogen |
CatNo: Ag17448 Product name: Recombinant human IFIH1 protein Source: e coli.-derived, PET28a Tag: 6*His Domain: 1-310 aa of BC111750 Sequence: MSNGYSTDENFRYLISCFRARVKMYIQVEPVLDYLTFLPAEVKEQIQRTVATSGNMQAVELLLSTLEKGVWHLGWTREFVEALRRTGSPLAARYMNPELTDLPSPSFENAHDEYLQLLNLLQPTLVDKLLVRDVLDKCMEEELLTIEDRNRIAAAENNGNESGVRELLKRIVQKENWFSAFLNVLRQTGNNELVQELTGSDCSESNAEIENLSQVDGPQVEEQLLSTTVQPNLEKEVWGMENNSSESSFADSSVVSESDTSLAEGSVSCLDESLGHNSNMGSDSGTMGSDSDEENVAARASPEPELQLRP Predict reactive species |
| Full Name | interferon induced with helicase C domain 1 |
| Calculated Molecular Weight | 1025 aa, 117 kDa |
| Observed Molecular Weight | 117-125 kDa |
| GenBank Accession Number | BC111750 |
| Gene Symbol | IFIH1 |
| Gene ID (NCBI) | 64135 |
| RRID | AB_2882116 |
| Conjugate | Unconjugated |
| Form | Liquid |
| Purification Method | Protein G purification |
| UNIPROT ID | Q9BYX4 |
| Storage Buffer | PBS with 0.02% sodium azide and 50% glycerol, pH 7.3. |
| Storage Conditions | Store at -20°C. Stable for one year after shipment. Aliquoting is unnecessary for -20oC storage. 20ul sizes contain 0.1% BSA. |
Background Information
IFIH1 (Interferon-induced helicase C domain-containing protein 1) is a putative RNA helicase that is upregulated in response to treatment with IFNB or IFNB and MEZ. It is also named as MDA5 and RH116. Ectopic expression of MDA5 in melanoma cells resulted in reduced colony formation, suggesting an interaction of the CARD and apoptotic signal molecules. Functional analysis indicated that MDA5 is an RNA-dependent ATPase. IFIH1 has an apparent molecular mass of 117-130 kDa, and always other bands (70 kDa and 90 kDa) can be detected as cleaved products (PMID: 17267501).
Protocols
| Product Specific Protocols | |
|---|---|
| FC protocol for IFIH1/MDA5 antibody 66770-1-Ig | Download protocol |
| IHC protocol for IFIH1/MDA5 antibody 66770-1-Ig | Download protocol |
| WB protocol for IFIH1/MDA5 antibody 66770-1-Ig | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |
Publications
| Species | Application | Title |
|---|---|---|
Adv Sci (Weinh) ADAR1 as a Placental Innate Immune Rheostat Sustaining the Homeostatic Balance of Intrinsic Interferon Response at the Maternal-Fetal Interface | ||
Virol Sin PRRSV degrades MDA5 via dual autophagy receptors P62 and CCT2 to evade antiviral innate immunity
| ||
J Virol Encephalomyocarditis virus abrogates the IFN-β signaling pathway via its structural protein VP2. | ||
Vet Microbiol DDX56 antagonizes IFN-β production to enhance EMCV replication by inhibiting IRF3 nuclear translocation | ||
Virus Res Host protein, HSP90β, antagonizes IFN-β signaling pathway and facilitates the proliferation of encephalomyocarditis virus in vitro. | ||
Front Immunol Temporally integrated transcriptome analysis reveals ASFV pathology and host response dynamics |
Reviews
The reviews below have been submitted by verified Proteintech customers who received an incentive for providing their feedback.
FH RASHMI (Verified Customer) (07-26-2024) | Used this product, highly recommended
|