Tested Applications
| Positive FC detected in | human PBMCs |
Recommended dilution
| Application | Dilution |
|---|---|
| Flow Cytometry (FC) | FC : 0.2 ug per 10^6 cells in 100 μl suspension |
| This reagent has been tested for flow cytometric analysis. It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
Published Applications
| WB | See 1 publications below |
| IHC | See 2 publications below |
Product Information
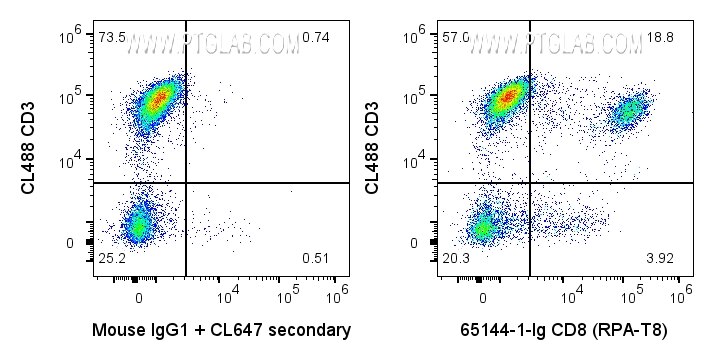
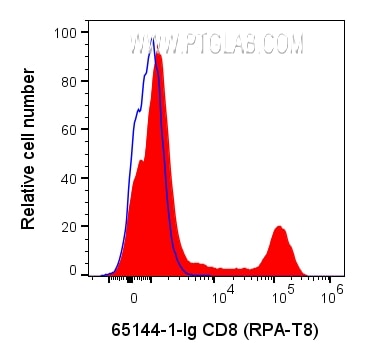
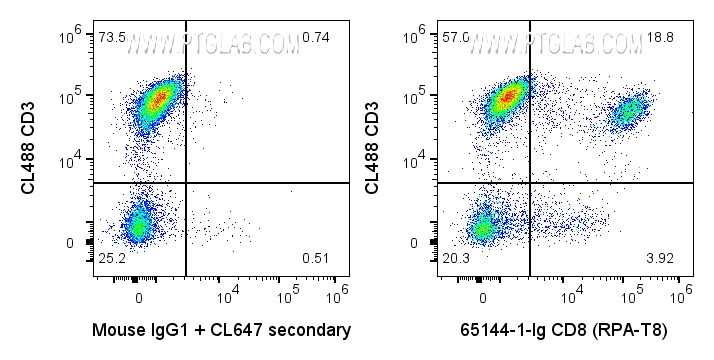
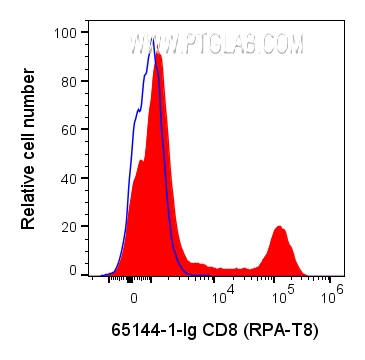
65144-1-Ig targets CD8a in FC applications and shows reactivity with human samples.
| Tested Reactivity | human |
| Cited Reactivity | human |
| Host / Isotype | Mouse / IgG1, kappa |
| Class | Monoclonal |
| Type | Antibody |
| Immunogen |
N/A Predict reactive species |
| Full Name | CD8a molecule |
| Calculated Molecular Weight | 235 aa, 26 kDa |
| GenBank Accession Number | BC025715 |
| Gene Symbol | CD8A |
| Gene ID (NCBI) | 925 |
| ENSEMBL Gene ID | ENSG00000153563 |
| RRID | AB_2918430 |
| Conjugate | Unconjugated |
| Form | Liquid |
| Purification Method | Affinity purification |
| UNIPROT ID | P01732 |
| Storage Buffer | PBS with 0.09% sodium azide, pH 7.3. |
| Storage Conditions | Store at 2-8°C. Stable for one year after shipment. |
Background Information
CD8 is a transmembrane glycoprotein composed of two disulfide-linked chains. It can be present as a homodimer of CD8α or as a heterodimer of CD8α and CD8β (PMID: 3264320; 8253791). CD8 is found on most thymocytes. The majority of class I-restricted T cells express mostly the CD8αβ heterodimer while CD8αα homodimers alone have been found on some gut intraepithelial T cells , on some T cell receptor (TCR) γδ T cells and on NK cells (PMID: 2111591; 1831127; 8420975). CD8 acts as a co-receptor that binds to MHC class-I and participates in cytotoxic T cell activation (PMID: 8499079). During T cell development, CD8 is required for positive selection of CD4-/CD8+ T cells (PMID: 1968084).
Protocols
| Product Specific Protocols | |
|---|---|
| FC protocol for CD8a antibody 65144-1-Ig | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |
Publications
| Species | Application | Title |
|---|---|---|
Cancers (Basel) Inhibition of DCLK1 with DCLK1-IN-1 Suppresses Renal Cell Carcinoma Invasion and Stemness and Promotes Cytotoxic T-Cell-Mediated Anti-Tumor Immunity. | ||
Front Oncol Comprehensive Characterization of Immune Landscape Based on Epithelial-Mesenchymal Transition Signature in OSCC: Implication for Prognosis and Immunotherapy. | ||
Front Oncol Typing and modeling of hepatocellular carcinoma based on disulfidptosis-related amino acid metabolism genes for predicting prognosis and guiding individualized treatment | ||
J Virol Combinational Deletions of MGF360-9L and MGF505-7R Attenuated Highly Virulent African Swine Fever Virus and Conferred Protection against Homologous Challenge. |