Tested Applications
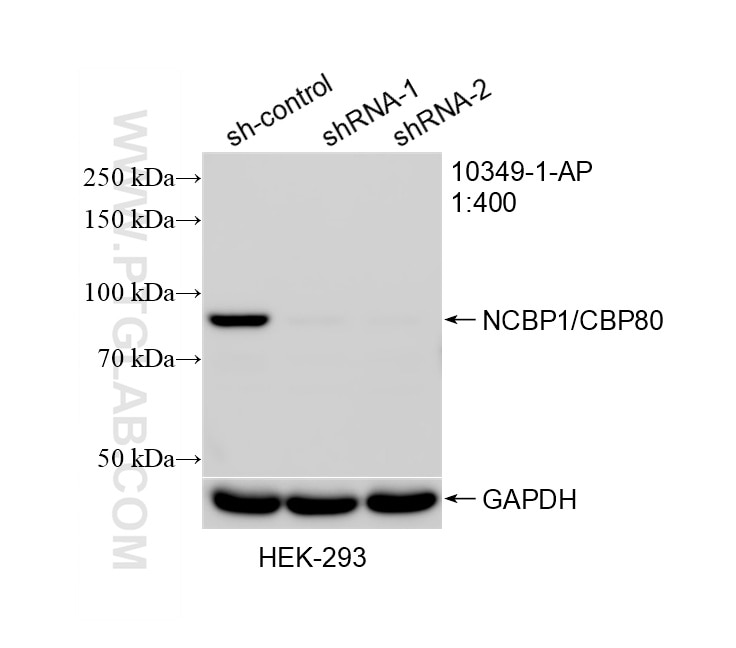
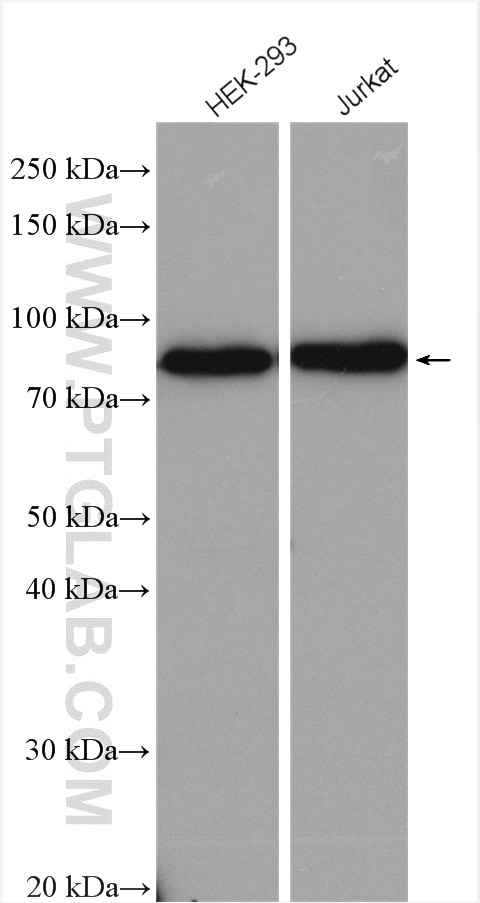
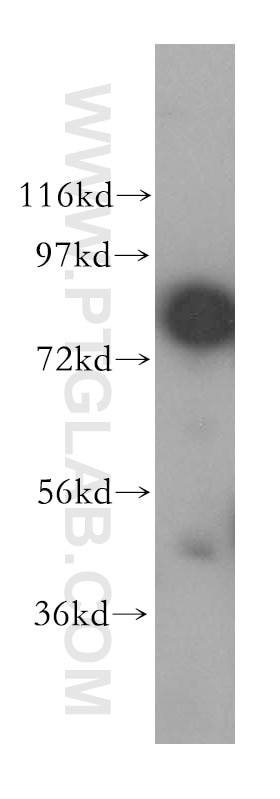
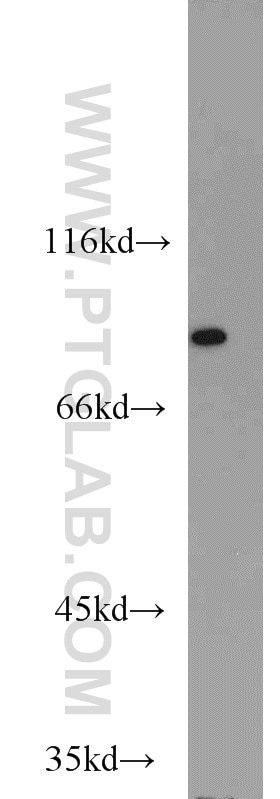
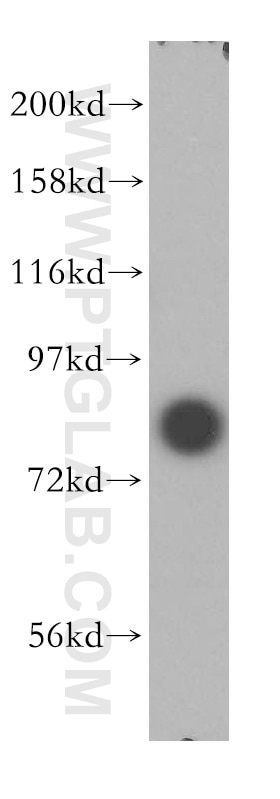
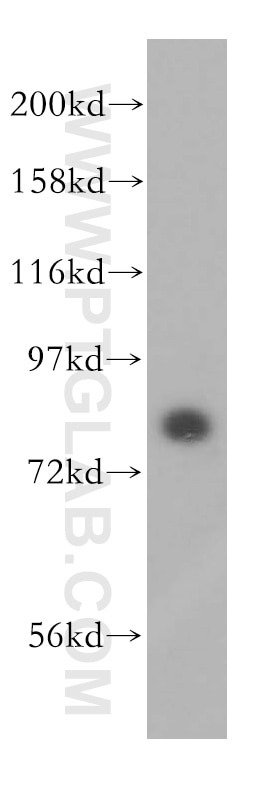
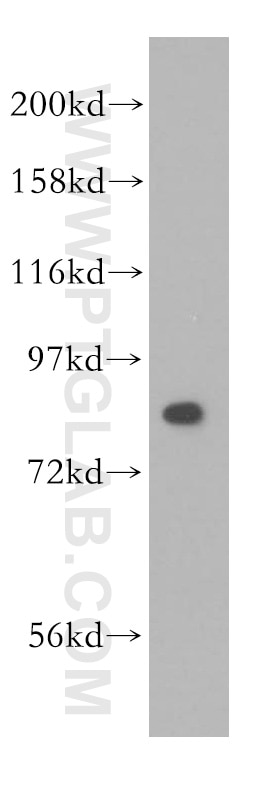
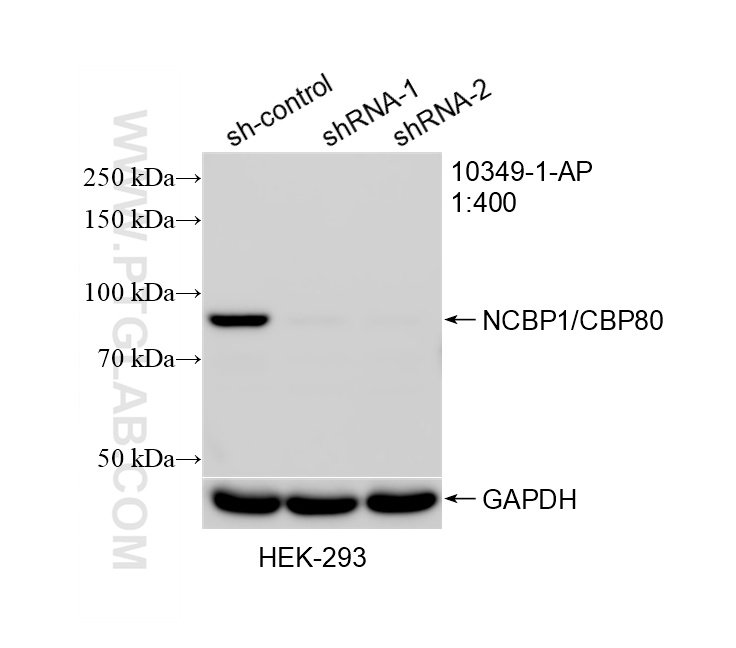
| Positive WB detected in | HEK-293 cells, HeLa cells, human brain tissue, Jurkat cells |
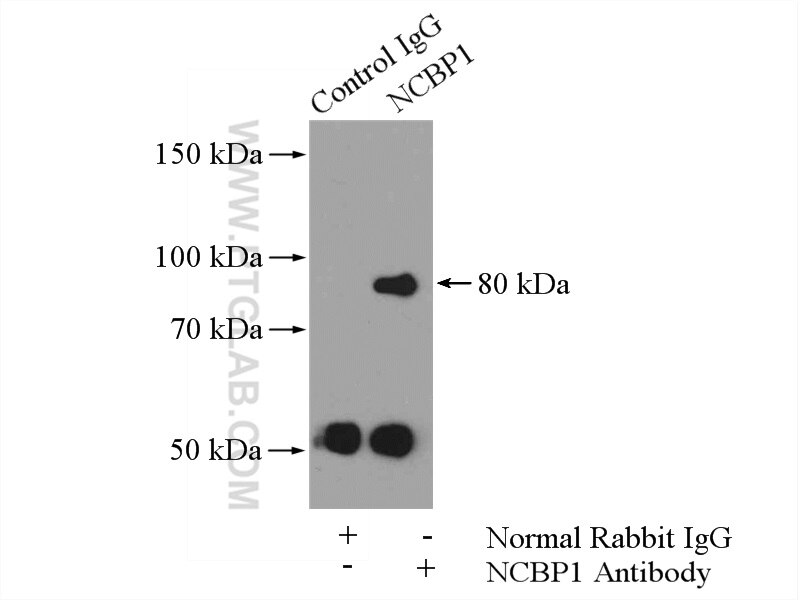
| Positive IP detected in | HeLa cells |
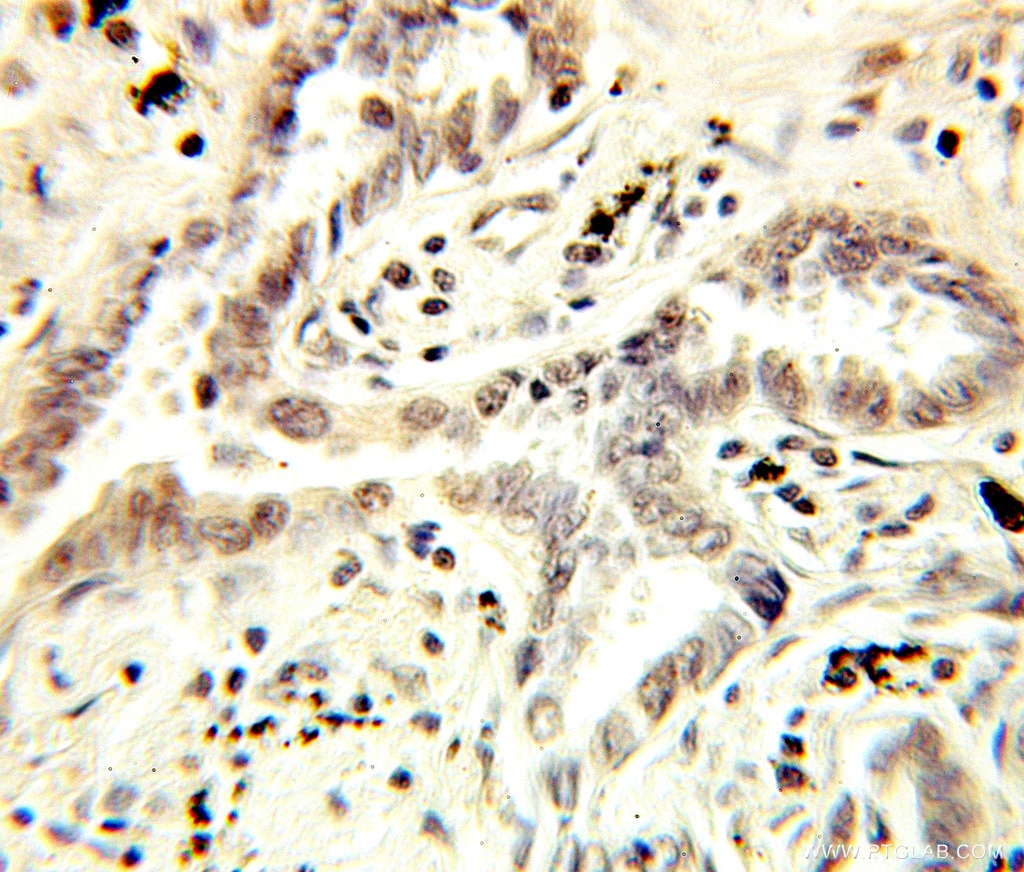
| Positive IHC detected in | human lung cancer tissue, human kidney tissue Note: suggested antigen retrieval with TE buffer pH 9.0; (*) Alternatively, antigen retrieval may be performed with citrate buffer pH 6.0 |
Recommended dilution
| Application | Dilution |
|---|---|
| Western Blot (WB) | WB : 1:500-1:2000 |
| Immunoprecipitation (IP) | IP : 0.5-4.0 ug for 1.0-3.0 mg of total protein lysate |
| Immunohistochemistry (IHC) | IHC : 1:20-1:200 |
| It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
Published Applications
| KD/KO | See 2 publications below |
| WB | See 12 publications below |
| IHC | See 1 publications below |
| IF | See 1 publications below |
| IP | See 3 publications below |
Product Information
10349-1-AP targets NCBP1/CBP80 in WB, IHC, IF, IP, ELISA applications and shows reactivity with human, mouse samples.
| Tested Reactivity | human, mouse |
| Cited Reactivity | human, mouse, monkey |
| Host / Isotype | Rabbit / IgG |
| Class | Polyclonal |
| Type | Antibody |
| Immunogen |
CatNo: Ag0496 Product name: Recombinant human CBP80 protein Source: e coli.-derived, PGEX-4T Tag: GST Domain: 46-350 aa of BC001450 Sequence: LESNLEGLAGVLEADLPNYKSKILRLLCTVARLLPEKLTIYTTLVGLLNARNYNFGGEFVEAMIRQLKESLKANNYNEAVYLVRFLSDLVNCHVIAAPSMVAMFENFVSVTQEEDVPQVRRDWYVYAFLSSLPWVGKELYEKKDAEMDRIFANTESYLKRRQKTHVPMLQVWTADKPHPQEEYLDCLWAQIQKLKKDRWQERHILRPYLAFDSILCEALQHNLPPFTPPPHTEDSVYPMPRVIFRMFDYTDDPEGPVMPGSHSVERFVIEENLHCIIKSHWKERKTCAAQLVSYPGKNKIPLNYH Predict reactive species |
| Full Name | nuclear cap binding protein subunit 1, 80kDa |
| Calculated Molecular Weight | 92 kDa |
| Observed Molecular Weight | 80 kDa |
| GenBank Accession Number | BC001450 |
| Gene Symbol | NCBP1 |
| Gene ID (NCBI) | 4686 |
| RRID | AB_2150127 |
| Conjugate | Unconjugated |
| Form | Liquid |
| Purification Method | Antigen affinity purification |
| UNIPROT ID | Q09161 |
| Storage Buffer | PBS with 0.02% sodium azide and 50% glycerol, pH 7.3. |
| Storage Conditions | Store at -20°C. Stable for one year after shipment. Aliquoting is unnecessary for -20oC storage. 20ul sizes contain 0.1% BSA. |
Background Information
Nuclear cap binding protein subunit 1, 80kDa (NCBP1, synonyms: NCBP, CBP80) is a subunit of a cap-binding protein complex (CBC) that consists of a CBP20 subunit, which binds the cap, and a NCBP1 subunit, which ensures high-affinity cap binding. The CBC complex may play a role in pre-mRNA recognition and splicing.
Protocols
| Product Specific Protocols | |
|---|---|
| IHC protocol for NCBP1/CBP80 antibody 10349-1-AP | Download protocol |
| IP protocol for NCBP1/CBP80 antibody 10349-1-AP | Download protocol |
| WB protocol for NCBP1/CBP80 antibody 10349-1-AP | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |
Publications
| Species | Application | Title |
|---|---|---|
Cell RNA-Methylation-Dependent RNA Processing Controls the Speed of the Circadian Clock.
| ||
Mol Cell RNA Pol II preferentially regulates ribosomal protein expression by trapping disassociated subunits | ||
Mol Cell Efficiency of the pioneer round of translation affects the cellular site of nonsense-mediated mRNA decay. | ||
Genes Dev SMG-8 and SMG-9, two novel subunits of the SMG-1 complex, regulate remodeling of the mRNA surveillance complex during nonsense-mediated mRNA decay. | ||
Nucleic Acids Res SUMOylation of the m6A-RNA methyltransferase METTL3 modulates its function. | ||
Sci Signal AAA+ proteins RUVBL1 and RUVBL2 coordinate PIKK activity and function in nonsense-mediated mRNA decay. |